Feral
RSP 10890
General Information
- Sample Name
- Merino_S_10_CSU
- Accession Date
- May 3, 2017
- Reported Plant Sex
- Female
- Report Type
- StrainSEEK v2 3.2Mb
- DNA Extracted From
- Unknown
The strain rarity visualization shows how distant the strain is from the other cultivars in the Kannapedia database. The y-axis represents genetic distance, getting farther as you go up. The width of the visualization at any position along the y-axis shows how many strains there are in the database at that genetic distance. So, a common strain will have a more bottom-heavy shape, while uncommon and rare cultivars will have a visualization that is generally shifted towards the top.
Chemical Information
Cannabinoid and terpenoid information provided by the grower.
Cannabinoids
No information provided.
Terpenoids
No information provided.
Genetic Information
- Plant Type
- Type III
File Downloads
The bell curve in the heterozygosity visualization shows the distribution of heterozygosity levels for cannabis cultivars in the Kannapedia database. The green line shows where this particular strain fits within the distribution. Heterozygosity is associated with heterosis (aka hybrid vigor) but also leads to the production of more variable offspring. When plants have two genetically different parents, heterozygosity levels will be higher than if it has been inbred or backcrossed repeatedly.
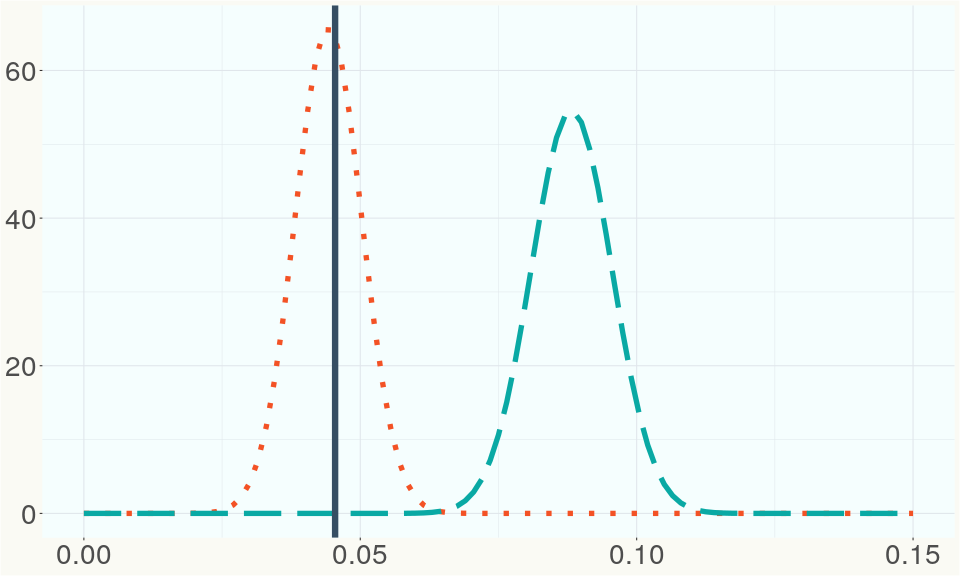
The ratio of reads mapped to Y-contigs to reads mapped to the whole Cannabis genome (Y-ratios) has been demonstrated to be strongly correlated with plant sex typing. This plot shows the distribution of Y-ratios for all samples in our database which were sequenced with the same method (panel or WGS) as this sample and where this sample falls in the distribution.
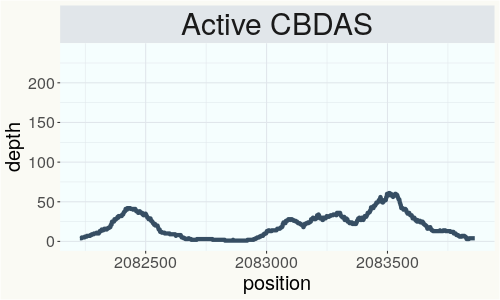
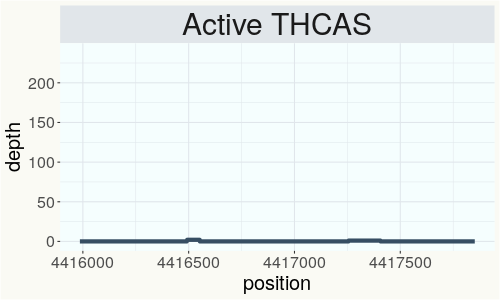
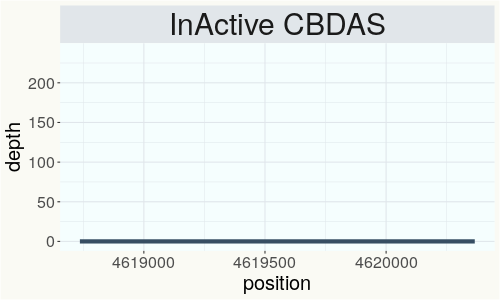
This chart represents the Illumina sequence coverage over the Bt/Bd allele. These are the three regions in the cannabis genome that impact THCA, CBDA, CBGA production. Coverage over the Active CBDAS gene is highly correlated with Type II and Type III plants as described by Etienne de Meijer. Coverage over the THCA gene is highly correlated with Type I and Type II plants but is anti-correlated with Type III plants. Type I plants require coverage over the inactive CBDA loci and no coverage over the Active CBDA gene. Lack of coverage over the Active CBDA and Active THCA allele are presumed to be Type IV plants (CBGA dominant). While deletions of entire THCAS and CBDAS genes are the most common Bt:Bd alleles observed, it is possible to have plants with these genes where functional expression of the enzyme is disrupted by deactivating point mutations (Kojoma et al. 2006).
This chart represents the Illumina sequence coverage over the CBCA synthase gene.
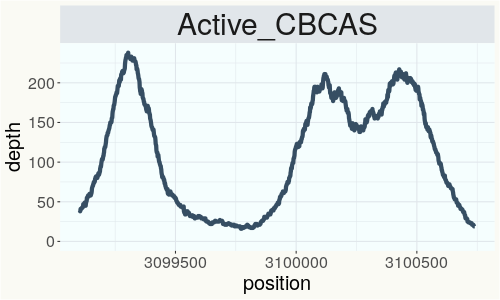
Variants (THCAS, CBDAS, and CBCAS)
Variants (Select Genes of Interest)
| GPPs1 | c.160A>G | p.Lys54Glu | missense variant | moderate | contig676 | 168409 | A/G |
|
| PKSG-4a | c.37C>G | p.Gln13Glu | missense variant | moderate | contig700 | 1936734 | C/G |
|
| PKSG-4a | c.493G>A | p.Gly165Ser | missense variant | moderate | contig700 | 1937904 | G/A |
|
| PKSG-4a |
c.1188_1190d |
p.Asn397del | disruptive inframe deletion | moderate | contig700 | 1938598 | CCAA/C |
|
| PKSG-4a | c.1192T>A | p.Tyr398Asn | missense variant | moderate | contig700 | 1938603 | T/A |
|
| PKSG-2a | c.1117A>G | p.Ile373Val | missense variant | moderate | contig700 | 1944273 | T/C | |
| PKSG-2a | c.948T>G | p.Asp316Glu | missense variant | moderate | contig700 | 1944442 | A/C |
|
| PKSG-2a | c.945T>G | p.Ser315Arg | missense variant | moderate | contig700 | 1944445 | A/C |
|
| PKSG-2a | c.944G>A | p.Ser315Asn | missense variant | moderate | contig700 | 1944446 | C/T |
|
| PKSG-2a | c.934C>G | p.His312Asp | missense variant | moderate | contig700 | 1944456 | G/C |
|
| PKSG-2b | c.1117A>G | p.Ile373Val | missense variant | moderate | contig700 | 1950521 | T/C | |
| PKSG-2b | c.67A>T | p.Ile23Phe | missense variant | moderate | contig700 | 1951815 | T/A | |
| PKSG-4b | c.544G>T | p.Gly182Trp | missense variant | moderate | contig700 | 2721129 | C/A |
|
| PKSG-4b | c.489delT | p.Phe163fs | frameshift variant | high | contig700 | 2721183 | CA/C | |
| PKSG-4b |
c.353_354ins |
p.Gly119fs | frameshift variant | high | contig700 | 2721319 | T/TGG |
|
| PKSG-4b | c.338G>A | p.Gly113Glu | missense variant | moderate | contig700 | 2721335 | C/T |
|
| DXR-2 | c.1319T>C | p.Ile440Thr | missense variant | moderate | contig380 | 285250 | A/G |
|
| aPT4 | c.80A>G | p.Lys27Arg | missense variant | moderate | contig121 | 2828736 | A/G |
|
| aPT4 | c.97T>C | p.Tyr33His | missense variant | moderate | contig121 | 2828753 | T/C |
|
| aPT4 | c.153A>C | p.Lys51Asn | missense variant | moderate | contig121 | 2828809 | A/C |
|
| aPT4 |
c.235_236del |
p.Val79fs | frameshift variant | high | contig121 | 2829030 | ATG/A |
|
| aPT4 | c.238delT | p.Ser80fs | frameshift variant | high | contig121 | 2829034 | AT/A |
|
| aPT4 | c.1168T>C | p.Tyr390His | missense variant | moderate | contig121 | 2833503 | T/C |
|
| aPT1 |
c.95_97delGT |
p.Cys32del | disruptive inframe deletion | moderate | contig121 | 2835800 | ATGT/A | |
| aPT1 | c.160A>C | p.Thr54Pro | missense variant | moderate | contig121 | 2835867 | A/C | |
| aPT1 | c.406A>G | p.Ile136Val | missense variant | moderate | contig121 | 2839605 | A/G | |
| aPT1 | c.670T>A | p.Ser224Thr | missense variant | moderate | contig121 | 2840278 | T/A |
|
| aPT1 | c.727G>T | p.Glu243* | stop gained | high | contig121 | 2841362 | G/T | |
| aPT1 | c.864C>G | p.Asn288Lys | missense variant | moderate | contig121 | 2842407 | C/G | |
| HDS-2 |
c.82_93delGT |
p.Val28_Thr3 |
conservative inframe deletion | moderate | contig95 | 1989748 |
CGTAACCGGAAC |
|
Nearest genetic relatives (All Samples)
- 0.166 Feral (RSP10891)
- 0.166 Feral (RSP10892)
- 0.179 Feral (RSP11205)
- 0.224 Carmagnola (RSP10978)
- 0.225 Carmagnola (RSP10982)
- 0.227 Carmagnola (RSP10976)
- 0.228 Carmagnola (RSP10979)
- 0.232 Carmagnola (RSP11039)
- 0.234 Carmagnola (RSP10980)
- 0.235 Tisza (RSP11044)
- 0.236 Carmagnola (RSP11037)
- 0.236 KYRG-151 (RSP11052)
- 0.236 Carmagnola (RSP11202)
- 0.237 Carmagnola (RSP10655)
- 0.240 VIR 483 (SRR14708238)
- 0.240 Carmagnola (RSP10977)
- 0.242 Feral (RSP11206)
- 0.243 Tisza (RSP11045)
- 0.244 Ivory (RSP10668)
- 0.244 Tisza (RSP10659)
Most genetically distant strains (All Samples)
- 0.451 Cherry Blossom (RSP11300)
- 0.443 Cherry Blossom (RSP11318)
- 0.438 QQD2 (RSP11450)
- 0.438 New York City Deisel (RSP11225)
- 0.438 Chem 91 (RSP11185)
- 0.436 Queen Dream CBG (RSP11287)
- 0.435 Escape Velocity (RSP11165)
- 0.433 Cherry Blossom (RSP11301)
- 0.432 Cherry Blossom (RSP11331)
- 0.432 Cherry Blossom (RSP11312)
- 0.431 Queen Dream (RSP11278)
- 0.430 UP Wendigo (RSP11261)
- 0.429 Northern Skunk (RSP11456)
- 0.428 JL x NSPM1 4 (RSP11482)
- 0.428 Cherry Blossom (RSP11323)
- 0.428 Up Sunset (RSP11256)
- 0.427 QLE1 (RSP11451)
- 0.427 GMO x Garlic Breath (RSP12507)
- 0.427 Big Bud (RSP11221)
- 0.427 East Coast Sour Diesel (RSP10243)
Nearest genetic relative in Phylos dataset
- Overlapping SNPs:
- 105
- Concordance:
- 65
Nearest genetic relative in Lynch dataset
- Overlapping SNPs:
- 3
- Concordance:
- 3
Blockchain Registration Information
- Transaction ID
-
4dd249199888858d
444a2d66126244db e82126ce81251b34 351365124bb10313 - Stamping Certificate
- Download PDF (854.5 KB)
- SHASUM Hash
-
105f22ccf92e1fd13af9b3aec07c1fc9 4115b99333826ccb 3b0d5f684392a58d