Durban Poison
RSP 11226
Grower: Los Suenos Farms
General Information
- Sample Name
- Pheno I
- Accession Date
- July 7, 2019
- Reported Plant Sex
- Female
- Report Type
- StrainSEEK v2 3.2Mb
- DNA Extracted From
- Stem
The strain rarity visualization shows how distant the strain is from the other cultivars in the Kannapedia database. The y-axis represents genetic distance, getting farther as you go up. The width of the visualization at any position along the y-axis shows how many strains there are in the database at that genetic distance. So, a common strain will have a more bottom-heavy shape, while uncommon and rare cultivars will have a visualization that is generally shifted towards the top.
Chemical Information
Cannabinoid and terpenoid information provided by the grower.
Cannabinoids
No information provided.
Terpenoids
No information provided.
Genetic Information
- Plant Type
- Type I
File Downloads
The bell curve in the heterozygosity visualization shows the distribution of heterozygosity levels for cannabis cultivars in the Kannapedia database. The green line shows where this particular strain fits within the distribution. Heterozygosity is associated with heterosis (aka hybrid vigor) but also leads to the production of more variable offspring. When plants have two genetically different parents, heterozygosity levels will be higher than if it has been inbred or backcrossed repeatedly.
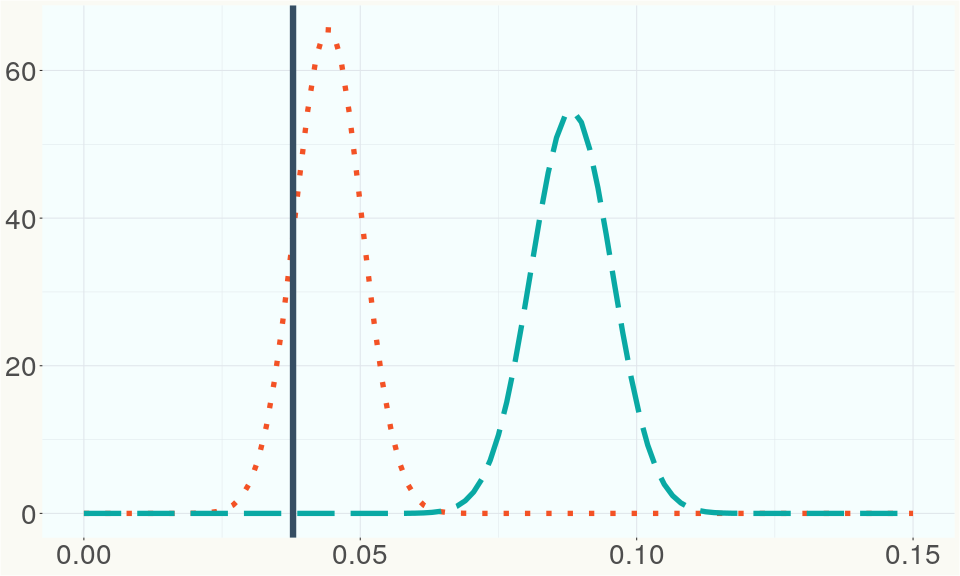
The ratio of reads mapped to Y-contigs to reads mapped to the whole Cannabis genome (Y-ratios) has been demonstrated to be strongly correlated with plant sex typing. This plot shows the distribution of Y-ratios for all samples in our database which were sequenced with the same method (panel or WGS) as this sample and where this sample falls in the distribution.
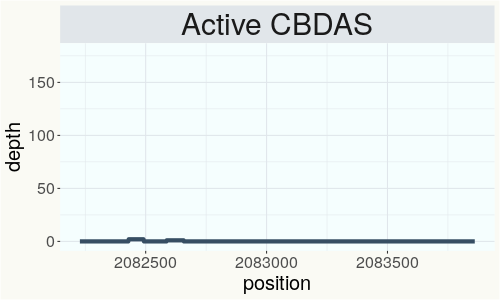
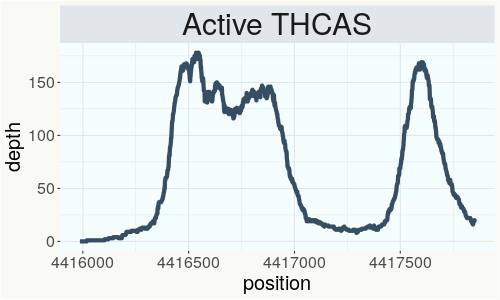
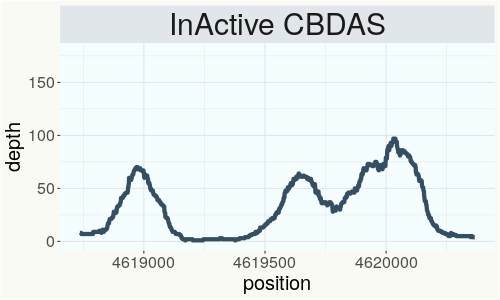
This chart represents the Illumina sequence coverage over the Bt/Bd allele. These are the three regions in the cannabis genome that impact THCA, CBDA, CBGA production. Coverage over the Active CBDAS gene is highly correlated with Type II and Type III plants as described by Etienne de Meijer. Coverage over the THCA gene is highly correlated with Type I and Type II plants but is anti-correlated with Type III plants. Type I plants require coverage over the inactive CBDA loci and no coverage over the Active CBDA gene. Lack of coverage over the Active CBDA and Active THCA allele are presumed to be Type IV plants (CBGA dominant). While deletions of entire THCAS and CBDAS genes are the most common Bt:Bd alleles observed, it is possible to have plants with these genes where functional expression of the enzyme is disrupted by deactivating point mutations (Kojoma et al. 2006).
This chart represents the Illumina sequence coverage over the CBCA synthase gene.
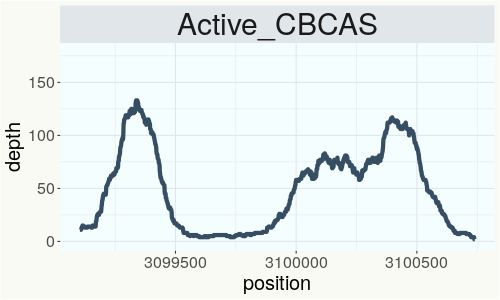
Variants (THCAS, CBDAS, and CBCAS)
Variants (Select Genes of Interest)
| PKSG-2a | c.774G>A | p.Met258Ile | missense variant | moderate | contig700 | 1944616 | C/T |
|
| PKSG-2a | c.67T>A | p.Phe23Ile | missense variant | moderate | contig700 | 1945567 | A/T | |
| PKSG-2a | c.31A>T | p.Thr11Ser | missense variant | moderate | contig700 | 1945603 | T/A | |
| PKSG-2b | c.1152T>A | p.Asn384Lys | missense variant | moderate | contig700 | 1950486 | A/T | |
| PKSG-2b | c.1132C>G | p.Leu378Val | missense variant | moderate | contig700 | 1950506 | G/C |
|
| PKSG-2b | c.1117A>G | p.Ile373Val | missense variant | moderate | contig700 | 1950521 | T/C | |
| PKSG-2b | c.1007T>C | p.Val336Ala | missense variant | moderate | contig700 | 1950631 | A/G |
|
| PKSG-2b | c.31A>T | p.Thr11Ser | missense variant | moderate | contig700 | 1951851 | T/A | |
| PKSG-4b | c.496A>G | p.Lys166Glu | missense variant | moderate | contig700 | 2721177 | T/C | |
| PKSG-4b | c.489delT | p.Phe163fs | frameshift variant | high | contig700 | 2721183 | CA/C | |
| PKSG-4b | c.485A>G | p.Lys162Arg | missense variant | moderate | contig700 | 2721188 | T/C | |
| PKSG-4b | c.431T>G | p.Val144Gly | missense variant | moderate | contig700 | 2721242 | A/C | |
| PKSG-4b | c.419A>G | p.Asp140Gly | missense variant | moderate | contig700 | 2721254 | T/C | |
| PKSG-4b |
c.352_355del |
p.Thr118fs | frameshift variant | high | contig700 | 2721317 | CCTGT/C |
|
| PKSG-4b | c.323A>G | p.Glu108Gly | missense variant | moderate | contig700 | 2721350 | T/C |
|
| DXR-2 | c.1319T>C | p.Ile440Thr | missense variant | moderate | contig380 | 285250 | A/G |
|
| FAD2-2 | c.161T>A | p.Leu54His | missense variant | moderate | contig83 | 1803208 | A/T |
|
| aPT4 | c.35A>C | p.Gln12Pro | missense variant | moderate | contig121 | 2828691 | A/C |
|
| aPT4 | c.97T>C | p.Tyr33His | missense variant | moderate | contig121 | 2828753 | T/C |
|
| aPT4 | c.153A>C | p.Lys51Asn | missense variant | moderate | contig121 | 2828809 | A/C |
|
| aPT4 | c.216A>T | p.Lys72Asn | missense variant | moderate | contig121 | 2828872 | A/T |
|
| aPT4 | c.775delT | p.Tyr259fs | frameshift variant | high | contig121 | 2831380 | AT/A |
|
| aPT4 | c.1168T>C | p.Tyr390His | missense variant | moderate | contig121 | 2833503 | T/C |
|
| aPT1 | c.406A>G | p.Ile136Val | missense variant | moderate | contig121 | 2839605 | A/G | |
| aPT1 | c.629C>T | p.Thr210Ile | missense variant | moderate | contig121 | 2840237 | C/T | |
| HDS-2 |
c.82_93delGT |
p.Val28_Thr3 |
conservative inframe deletion | moderate | contig95 | 1989748 |
CGTAACCGGAAC |
|
| HDS-2 | c.127T>G | p.Ser43Ala | missense variant | moderate | contig95 | 1989794 | T/G |
|
| HDS-1 |
c.-108+1_-10 |
splice donor variant & intron variant | high | contig1891 | 889975 | A/AC |
|
Nearest genetic relatives (All Samples)
- 0.002 Durban Poison (RSP10998)
- 0.004 Durban Poison #1 (RSP11013)
- 0.006 Durban Poison (RSP11014)
- 0.006 Durban Poison #1 (RSP10996)
- 0.159 Black Jack (RSP10603)
- 0.165 BLACK JACK (RSP11346)
- 0.166 RKM-2018-025 (RSP11117)
- 0.166 RKM-2018-016 (RSP11108)
- 0.171 Domnesia (RSP11184)
- 0.185 Electra (RSP11366)
- 0.190 Lift (RSP11378)
- 0.192 Gold Cracker (RSP11048)
- 0.194 Saint Jack (RSP11179)
- 0.197 Cheese (RSP10460)
- 0.199 RKM-2018-027 (RSP11119)
- 0.205 UnObtanium (RSP11611)
- 0.209 Joy (RSP11380)
- 0.209 Doug s Varin (RSP11243)
- 0.212 Strawnana (RSP12094)
- 0.213 Liberty Haze (RSP11000)
Most genetically distant strains (All Samples)
- 0.442 Cherry Blossom (RSP11323)
- 0.428 80E (RSP11213)
- 0.415 Cherry Blossom (RSP11318)
- 0.409 Cherry Blossom (RSP11314)
- 0.407 80E (RSP11212)
- 0.404 Cherry Blossom (RSP11317)
- 0.402 80E (RSP11211)
- 0.401 CS (RSP11208)
- 0.396 Kush Hemp E1 (RSP11128)
- 0.395 Cherry Blossom (RSP11333)
- 0.392 Chematonic -Cannatonic x Chemdawg- (RSP11394)
- 0.390 R1in136 (SRR14708227)
- 0.388 Cherry Blossom (RSP11311)
- 0.388 IUP3 (SRR14708256)
- 0.388 GMO x Garlic Breath (RSP12507)
- 0.387 IUP2 (SRR14708257)
- 0.385 Feral (RSP11206)
- 0.385 Carmagnola (RSP11037)
- 0.385 Feral (RSP11205)
- 0.384 ILM (RSP12623)
Nearest genetic relative in Phylos dataset
- Overlapping SNPs:
- 98
- Concordance:
- 95
Nearest genetic relative in Lynch dataset
- Overlapping SNPs:
- 11
- Concordance:
- 10
Blockchain Registration Information
- Transaction ID
-
498383fbfdc73e89
1f76cc0458de64dd c97adf777d204e58 787a17631d511111 - Stamping Certificate
- Download PDF (840.6 KB)
- SHASUM Hash
-
54fc8622ad6d7829b2f87aa812d8af89 6d2c12daeb9c1cdd df04f388adfb52ee