Cherry Sunrise
RSP 11633
Grower: Trilogene Seeds
General Information
The strain rarity visualization shows how distant the strain is from the other cultivars in the Kannapedia database. The y-axis represents genetic distance, getting farther as you go up. The width of the visualization at any position along the y-axis shows how many strains there are in the database at that genetic distance. So, a common strain will have a more bottom-heavy shape, while uncommon and rare cultivars will have a visualization that is generally shifted towards the top.
Chemical Information
Cannabinoid and terpenoid information provided by the grower.
Cannabinoids
No information provided.
Terpenoids
No information provided.
Genetic Information
- Plant Type
- Type III
File Downloads
This chart represents the Log-R Ratio (LRR) over variants in the region of the THCA synthase gene. A high correlation between the LRR and samples with a known deletion in THCA synthase indicates the THCA region is deleted and a low correlation indicates it is intact.
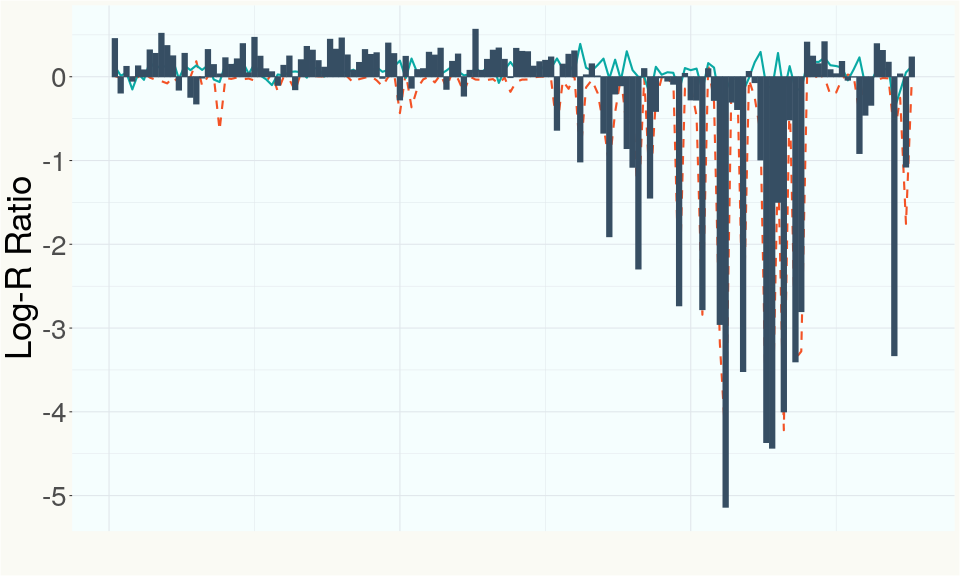
This chart represents the Log-R Ratio (LRR) over variants in the region of the CBDA synthase gene. A high correlation between the LRR and samples with a known deletion in CBDA synthase indicates the CBDA region is deleted and a low correlation indicates it is intact.
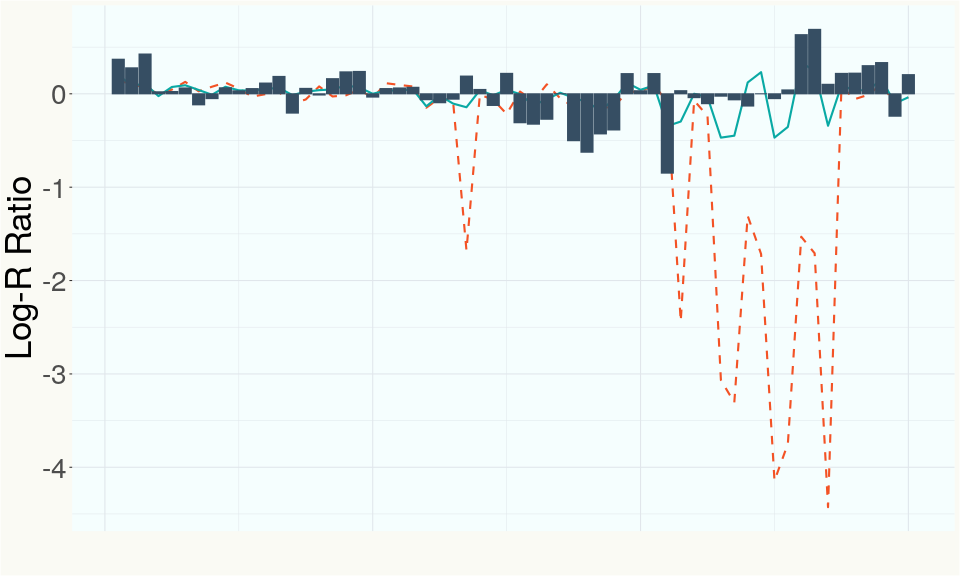
This chart represents the Log-R Ratio (LRR) over variants in the region of the CBCA synthase gene. A high correlation between the LRR and samples with a known deletion in CBCA synthase indicates the CBCA region is deleted and a low correlation indicates it is intact.
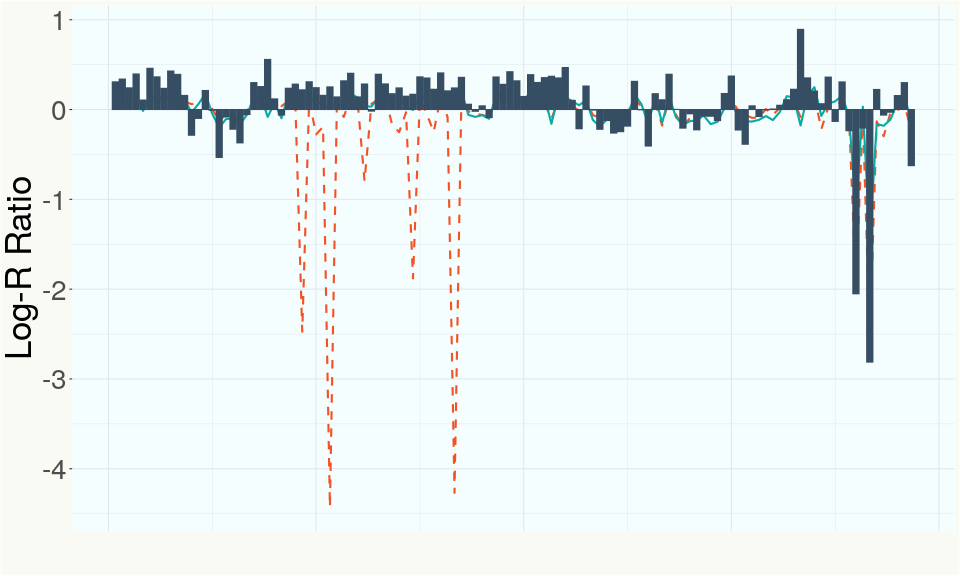
This chart represents the Log-R Ratio (LRR) over variants in the Y-contigs. A high correlation between the LRR and samples which are known Females indicates these Y-contigs are deleted in this sample and a low correlation indicates that the Y-contigs are not deleted and is likely Male.
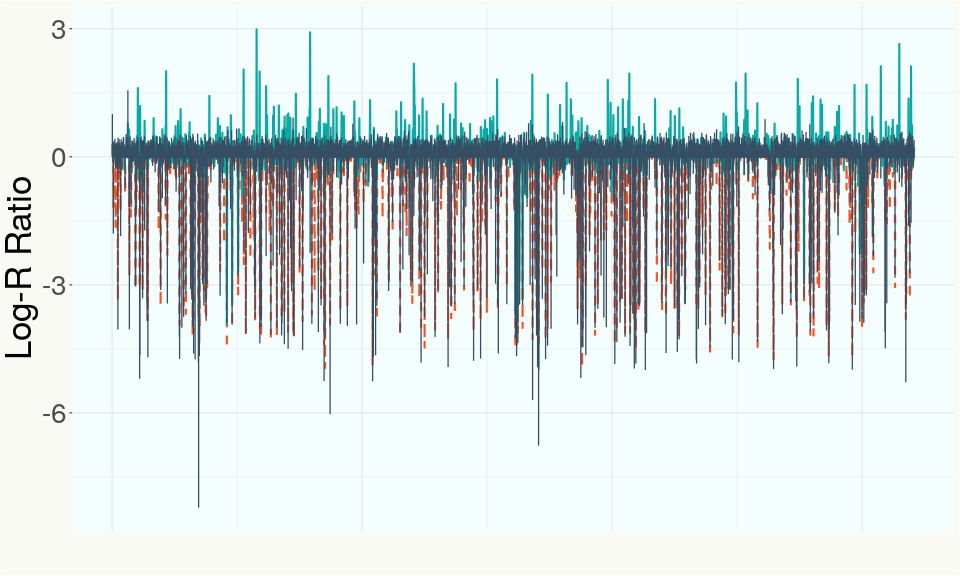
Summary of Deletions
THCAS
- Correlation:
- 0.95
- Call:
- deleted
CBDAS
- Correlation:
- -0.08
- Call:
- intact
CBCAS
- Correlation:
- 0.33
- Call:
- intact
Plant Sex
- Correlation:
- 0.93
- Call:
- female
Variants (THCAS, CBDAS, and CBCAS)
Variants (Select Genes of Interest)
| EMF1-2 | c.634G>C | p.Val212Leu | missense variant | moderate | contig885 | 734 | G/C | |
| EMF1-2 | c.710A>C | p.His237Pro | missense variant | moderate | contig885 | 810 | A/C | |
| EMF1-2 | c.1384A>C | p.Lys462Gln | missense variant | moderate | contig885 | 2270 | A/C | |
| PHL-2 | c.1057A>G | p.Arg353Gly | missense variant | moderate | contig2621 | 340335 | A/G | |
| PHL-2 | c.1096G>A | p.Ala366Thr | missense variant | moderate | contig2621 | 340374 | G/A | |
| PHL-2 | c.1540A>G | p.Thr514Ala | missense variant | moderate | contig2621 | 340818 | A/G | |
| PHL-2 | c.2756A>C | p.Glu919Ala | missense variant | moderate | contig2621 | 342799 | A/C | |
| PHL-2 | c.2783G>A | p.Ser928Asn | missense variant | moderate | contig2621 | 342826 | G/A | |
| PHL-2 | c.3002A>G | p.Tyr1001Cys | missense variant | moderate | contig2621 | 343045 | A/G | |
| PHL-2 | c.3027G>T | p.Lys1009Asn | missense variant | moderate | contig2621 | 343070 | G/T | |
| PHL-2 | c.3033T>G | p.Cys1011Trp | missense variant | moderate | contig2621 | 343076 | T/G | |
| PHL-2 | c.3209A>G | p.Gln1070Arg | missense variant | moderate | contig2621 | 343252 | A/G | |
| PKSG-4a | c.1000T>C | p.Tyr334His | missense variant | moderate | contig700 | 1938411 | T/C | |
| PKSG-2a | c.1152T>A | p.Asn384Lys | missense variant | moderate | contig700 | 1944238 | A/T | |
| PKSG-2a | c.1117A>G | p.Ile373Val | missense variant | moderate | contig700 | 1944273 | T/C | |
| PKSG-2a | c.224A>G | p.Lys75Arg | missense variant | moderate | contig700 | 1945166 | T/C | |
| PKSG-2a | c.67T>A | p.Phe23Ile | missense variant | moderate | contig700 | 1945567 | A/T | |
| PKSG-2a | c.31A>T | p.Thr11Ser | missense variant | moderate | contig700 | 1945603 | T/A | |
| PKSG-2b | c.1152T>A | p.Asn384Lys | missense variant | moderate | contig700 | 1950486 | A/T | |
| PKSG-2b | c.1117A>G | p.Ile373Val | missense variant | moderate | contig700 | 1950521 | T/C | |
| PKSG-2b | c.774G>A | p.Met258Ile | missense variant | moderate | contig700 | 1950864 | C/T | |
| PKSG-2b | c.224A>G | p.Lys75Arg | missense variant | moderate | contig700 | 1951414 | T/C | |
| PKSG-2b | c.167C>G | p.Thr56Ser | missense variant | moderate | contig700 | 1951471 | G/C | |
| PKSG-2b | c.31A>T | p.Thr11Ser | missense variant | moderate | contig700 | 1951851 | T/A | |
| PKSG-4b | c.496A>G | p.Lys166Glu | missense variant | moderate | contig700 | 2721177 | T/C | |
| PKSG-4b | c.485A>G | p.Lys162Arg | missense variant | moderate | contig700 | 2721188 | T/C | |
| PKSG-4b | c.431T>G | p.Val144Gly | missense variant | moderate | contig700 | 2721242 | A/C | |
| PKSG-4b | c.316+2T>A | splice donor variant & intron variant | high | contig700 | 2723818 | A/T | ||
| ELF3 | c.574A>G | p.Asn192Asp | missense variant | moderate | contig97 | 242280 | A/G | |
| ELF3 | c.812G>C | p.Gly271Ala | missense variant | moderate | contig97 | 242518 | G/C | |
| ELF3 | c.1630A>G | p.Thr544Ala | missense variant | moderate | contig97 | 244461 | A/G | |
| ELF3 | c.1966C>G | p.Pro656Ala | missense variant | moderate | contig97 | 244797 | C/G | |
| ELF3 | c.2198delG | p.Arg733fs | frameshift variant | high | contig97 | 245028 | CG/C | |
| ELF3 | c.2198G>T | p.Arg733Leu | missense variant | moderate | contig97 | 245029 | G/T | |
| ELF3 | c.2216A>G | p.His739Arg | missense variant | moderate | contig97 | 245047 | A/G | |
| aPT1 | c.629C>T | p.Thr210Ile | missense variant | moderate | contig121 | 2840237 | C/T | |
| PHL-1 | c.2551A>G | p.Thr851Ala | missense variant | moderate | contig1439 | 1487246 | T/C | |
| PHL-1 | c.1387A>G | p.Thr463Ala | missense variant | moderate | contig1439 | 1489811 | T/C | |
| HDS-1 | c.136G>A | p.Val46Ile | missense variant | moderate | contig1891 | 889256 | C/T | |
| HDS-1 | c.56C>G | p.Ala19Gly | missense variant | moderate | contig1891 | 889336 | G/C | |
| HDS-1 | c.35G>A | p.Cys12Tyr | missense variant | moderate | contig1891 | 889357 | C/T | |
| PIE1-2 | c.6653A>G | p.Asn2218Ser | missense variant | moderate | contig1460 | 1184434 | T/C | |
| PIE1-2 | c.6636T>G | p.Asp2212Glu | missense variant | moderate | contig1460 | 1184451 | A/C | |
| PIE1-2 | c.5932A>G | p.Ile1978Val | missense variant | moderate | contig1460 | 1185552 | T/C | |
| PIE1-2 | c.5132T>C | p.Ile1711Thr | missense variant | moderate | contig1460 | 1186607 | A/G |
|
| PIE1-2 | c.2188G>A | p.Ala730Thr | missense variant | moderate | contig1460 | 1189852 | C/T | |
| PIE1-2 |
c.2083_2085d |
p.Val695del | conservative inframe deletion | moderate | contig1460 | 1189954 | GGAC/G | |
| PIE1-2 | c.2072A>G | p.His691Arg | missense variant | moderate | contig1460 | 1189968 | T/C | |
| PIE1-2 | c.1872T>A | p.Asp624Glu | missense variant | moderate | contig1460 | 1190252 | A/T | |
| PIE1-2 | c.1656T>G | p.Asn552Lys | missense variant | moderate | contig1460 | 1191574 | A/C | |
| PIE1-2 | c.1630G>C | p.Ala544Pro | missense variant | moderate | contig1460 | 1191600 | C/G | |
| PIE1-2 | c.1156T>G | p.Trp386Gly | missense variant | moderate | contig1460 | 1192242 | A/C | |
| PIE1-2 | c.1117C>G | p.Gln373Glu | missense variant | moderate | contig1460 | 1192281 | G/C | |
| PIE1-2 | c.1093G>A | p.Gly365Ser | missense variant | moderate | contig1460 | 1192305 | C/T | |
| PIE1-2 | c.982G>A | p.Glu328Lys | missense variant | moderate | contig1460 | 1192416 | C/T | |
| PIE1-2 | c.710C>T | p.Pro237Leu | missense variant | moderate | contig1460 | 1193804 | G/A | |
| PIE1-2 | c.706T>C | p.Tyr236His | missense variant | moderate | contig1460 | 1193808 | A/G | |
| PIE1-2 | c.637T>A | p.Ser213Thr | missense variant | moderate | contig1460 | 1194421 | A/T | |
| PIE1-2 | c.349C>T | p.Pro117Ser | missense variant & splice region variant | moderate | contig1460 | 1195017 | G/A |
|
| EMF2 | c.1772A>G | p.Gln591Arg | missense variant | moderate | contig954 | 3059929 | A/G | |
| EMF1-1 | c.470C>A | p.Ser157Tyr | missense variant | moderate | contig883 | 269959 | C/A | |
| EMF1-1 | c.605A>T | p.His202Leu | missense variant | moderate | contig883 | 270210 | A/T | |
| FLD | c.2981T>C | p.Met994Thr | missense variant | moderate | contig1450 | 2044012 | A/G | |
| FLD | c.2964C>A | p.Asp988Glu | missense variant | moderate | contig1450 | 2044029 | G/T | |
| FLD | c.2686G>A | p.Ala896Thr | missense variant | moderate | contig1450 | 2044848 | C/T | |
| FLD | c.2681T>C | p.Ile894Thr | missense variant | moderate | contig1450 | 2044853 | A/G | |
| FLD | c.125G>A | p.Ser42Asn | missense variant | moderate | contig1450 | 2047909 | C/T | |
| AAE1-3 | c.3G>A | p.Met1? | start lost | high | contig976 | 1084072 | C/T | |
| PIE1-1 | c.742T>A | p.Ser248Thr | missense variant | moderate | contig1225 | 2279320 | T/A | |
| PIE1-1 | c.773A>G | p.Asn258Ser | missense variant & splice region variant | moderate | contig1225 | 2279897 | A/G | |
| PIE1-1 | c.811T>C | p.Tyr271His | missense variant | moderate | contig1225 | 2279935 | T/C | |
| PIE1-1 | c.815C>T | p.Pro272Leu | missense variant | moderate | contig1225 | 2279939 | C/T | |
| PIE1-1 | c.1222C>G | p.Gln408Glu | missense variant | moderate | contig1225 | 2281482 | C/G | |
| PIE1-1 | c.1394A>G | p.Asp465Gly | missense variant | moderate | contig1225 | 2281654 | A/G | |
| PIE1-1 | c.1454T>C | p.Val485Ala | missense variant | moderate | contig1225 | 2281714 | T/C |
|
| PIE1-1 |
c.1548_1549i |
p.Gln516_Glu |
conservative inframe insertion | moderate | contig1225 | 2281807 | A/AGAT | |
| PIE1-1 | c.1758T>G | p.Asn586Lys | missense variant | moderate | contig1225 | 2282186 | T/G | |
| PIE1-1 | c.2174A>G | p.His725Arg | missense variant | moderate | contig1225 | 2283789 | A/G |
|
| PIE1-1 |
c.2185_2187d |
p.Val729del | conservative inframe deletion | moderate | contig1225 | 2283796 | AGTC/A |
|
| PIE1-1 | c.3869A>C | p.Asn1290Thr | missense variant | moderate | contig1225 | 2285484 | A/C | |
| PIE1-1 | c.5234T>C | p.Ile1745Thr | missense variant | moderate | contig1225 | 2287152 | T/C |
|
| PIE1-1 | c.6041T>C | p.Met2014Thr | missense variant | moderate | contig1225 | 2288214 | T/C |
|
| PIE1-1 | c.6738G>T | p.Glu2246Asp | missense variant | moderate | contig1225 | 2289303 | G/T |
|
| GGR | c.376G>C | p.Glu126Gln | missense variant | moderate | contig2282 | 549368 | G/C | |
| GGR | c.581C>T | p.Ala194Val | missense variant | moderate | contig2282 | 549573 | C/T |
Nearest genetic relatives (All Samples)
- 0.097 Lifter (RSP11365)
- 0.108 Calm (RSP11379)
- 0.115 Lift (RSP11378)
- 0.118 Electra (RSP11366)
- 0.121 JL Cross 78 (RSP11579)
- 0.131 JL Cross 84 (RSP11585)
- 0.133 WO DC (RSP11413)
- 0.134 Herijuana (RSP11181)
- 0.136 Aquawoman (RSP11634)
- 0.136 JL Cross 77 (RSP11578)
- 0.136 JL Cross 51 (RSP11552)
- 0.137 Domnesia (RSP11184)
- 0.137 Thank You Jerry (RSP11459)
- 0.137 Charlotte Dream (RSP11412)
- 0.137 JL Cross 35 (RSP11536)
- 0.138 Wedding Cake x MAC (RSP11464)
- 0.139 Cbot-2019-001 (RSP11129)
- 0.139 JL Cross 80 (RSP11581)
- 0.139 Sangria (RSP11631)
- 0.139 JL Cross 10 (RSP11511)
Most genetically distant strains (All Samples)
- 0.244 Feral (RSP11205)
- 0.241 Tiborszallasie (RSP11210)
- 0.240 CS (RSP11208)
- 0.238 Fedora 17 (RSP11203)
- 0.236 Carmaleonte (RSP11207)
- 0.234 Carmagnola USO 31 (RSP11204)
- 0.230 Feral (RSP11206)
- 0.228 80E (RSP11212)
- 0.224 80E (RSP11211)
- 0.223 80E (RSP11213)
- 0.222 Eletta Campana (RSP11209)
- 0.208 Arcata Trainwreck (RSP11176)
- 0.207 JL 3rd Gen Father (RSP11196)
- 0.206 Goomendaze (RSP11462)
- 0.203 JL 4th Gen 6 (RSP11200)
- 0.195 JL 4th Gen 1 (RSP11193)
- 0.195 JL Cross 33 (RSP11534)
- 0.191 Red Eye OG (RSP11190)
- 0.191 JL 4th Gen 2 (RSP11194)
- 0.186 Compost (RSP11602)
Nearest genetic relative in Phylos dataset
- Overlapping SNPs:
- 86
- Concordance:
- 53
Nearest genetic relative in Lynch dataset
- Overlapping SNPs:
- 2
- Concordance:
- 2
Blockchain Registration Information
- SHASUM Hash
-
fcb39f5f87abd8f4c31febb13102cf23 c3715e07e1663acb afdfa59fa68ab6a3