Jamaican Lion
RSP 12915
Grower: kevin McKernan
General Information
- Sample Name
- JL-Fem-Seed-Flower
- Accession Date
- September 14, 2023
- Reported Plant Sex
- Female
- Report Type
- Whole-Genome Sequencing
- Microbiome
- Krona Plot
The strain rarity visualization shows how distant the strain is from the other cultivars in the Kannapedia database. The y-axis represents genetic distance, getting farther as you go up. The width of the visualization at any position along the y-axis shows how many strains there are in the database at that genetic distance. So, a common strain will have a more bottom-heavy shape, while uncommon and rare cultivars will have a visualization that is generally shifted towards the top.
Chemical Information
Cannabinoid and terpenoid information provided by the grower.
Cannabinoids
No information provided.
Terpenoids
No information provided.
Genetic Information
- Plant Type
- Type II
File Downloads
The bell curve in the heterozygosity visualization shows the distribution of heterozygosity levels for cannabis cultivars in the Kannapedia database. The green line shows where this particular strain fits within the distribution. Heterozygosity is associated with heterosis (aka hybrid vigor) but also leads to the production of more variable offspring. When plants have two genetically different parents, heterozygosity levels will be higher than if it has been inbred or backcrossed repeatedly.
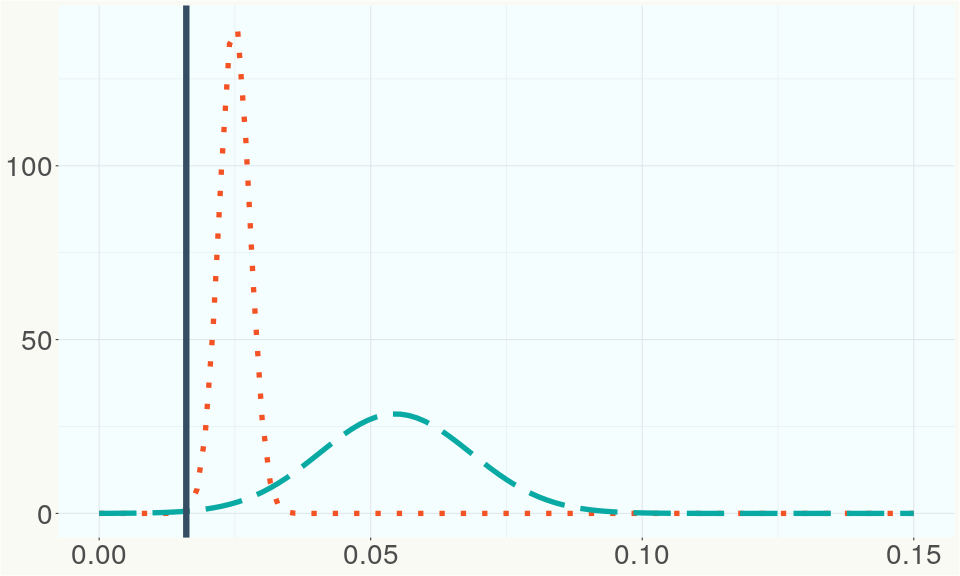
The ratio of reads mapped to Y-contigs to reads mapped to the whole Cannabis genome (Y-ratios) has been demonstrated to be strongly correlated with plant sex typing. This plot shows the distribution of Y-ratios for all samples in our database which were sequenced with the same method (panel or WGS) as this sample and where this sample falls in the distribution.
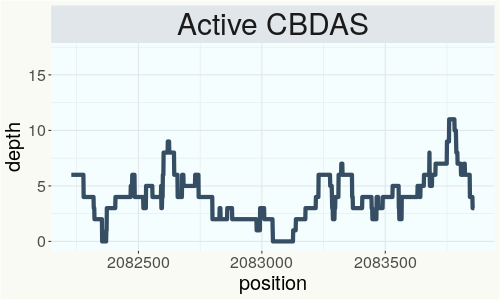
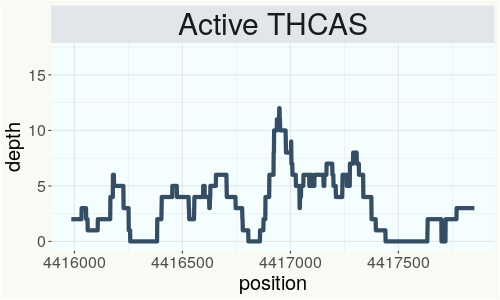
This chart represents the Illumina sequence coverage over the Bt/Bd allele. These are the three regions in the cannabis genome that impact THCA, CBDA, CBGA production. Coverage over the Active CBDAS gene is highly correlated with Type II and Type III plants as described by Etienne de Meijer. Coverage over the THCA gene is highly correlated with Type I and Type II plants but is anti-correlated with Type III plants. Type I plants require coverage over the inactive CBDA loci and no coverage over the Active CBDA gene. Lack of coverage over the Active CBDA and Active THCA allele are presumed to be Type IV plants (CBGA dominant). While deletions of entire THCAS and CBDAS genes are the most common Bt:Bd alleles observed, it is possible to have plants with these genes where functional expression of the enzyme is disrupted by deactivating point mutations (Kojoma et al. 2006).
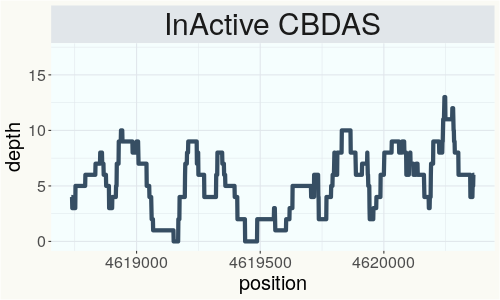
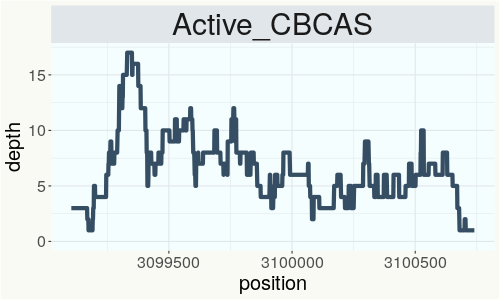
This chart represents the Illumina sequence coverage over the CBCA synthase gene.
Variants (THCAS, CBDAS, and CBCAS)
No variants to report
Variants (Select Genes of Interest)
| FAD2-2 | c.62C>T | p.Pro21Leu | missense variant | moderate | contig83 | 1803307 | G/A |
|
| ELF3 |
c.1803_1805d |
p.His601del | disruptive inframe deletion | moderate | contig97 | 244625 | ACAT/A |
|
| AAE1-2 | c.331A>G | p.Asn111Asp | missense variant | moderate | contig81 | 209293 | A/G |
|
| PHL-1 | c.1387A>G | p.Thr463Ala | missense variant | moderate | contig1439 | 1489811 | T/C | |
| AAE1-3 | c.181G>A | p.Val61Ile | missense variant | moderate | contig976 | 1083894 | C/T |
|
| AAE1-3 | c.167A>G | p.Glu56Gly | missense variant | moderate | contig976 | 1083908 | T/C |
|
| AAE1-3 | c.125A>G | p.Glu42Gly | missense variant | moderate | contig976 | 1083950 | T/C |
|
| AAE1-3 | c.79A>G | p.Thr27Ala | missense variant | moderate | contig976 | 1083996 | T/C |
|
| GGR | c.317C>T | p.Pro106Leu | missense variant | moderate | contig2282 | 549309 | C/T | |
| GGR | c.704A>T | p.His235Leu | missense variant | moderate | contig2282 | 549696 | A/T |
|
Nearest genetic relatives (All Samples)
- 0.051 JL yellow (RSP11075)
- 0.052 JL 3rd Gen Mother (RSP11214)
- 0.059 JL 3rd Gen Mother (RSP11197)
- 0.065 Jamaican Lion (RSP12917)
- 0.082 Jamaican Lion (RSP12916)
- 0.163 JL 4th Gen 1 (RSP11193)
- 0.175 JL 4th Gen 2 (RSP11194)
- 0.181 JL 4th Gen 5 (RSP11199)
- 0.186 JL 4th Gen 4 (RSP11198)
- 0.187 JL 4th Gen 3 (RSP11195)
- 0.199 JL 4th Gen 6 (RSP11200)
- 0.214 Original Jamaican Lions (RSP11127)
- 0.239 JL X NSPM1 12 (RSP11472)
- 0.241 JL 4th Gen 7 (RSP11153)
- 0.243 Jamaican Lion (RSP12920)
- 0.243 Jamaican Lion (RSP12919)
- 0.243 JL x NSPM1 4 (RSP11482)
- 0.246 Jamaican Lion (RSP12918)
- 0.271 JL Tent 1 yellow stake (RSP11488)
- 0.273 JL Tent 4 (RSP11491)
Nearest genetic relatives (Base Tree)
- 0.049 JL yellow (RSP11075)
- 0.310 RKM-2018-006 (RSP11097)
- 0.312 CST (RSP11002)
- 0.314 Sour Raspberry (RSP10551)
- 0.314 Blue Dream (RSP11033)
- 0.315 UP Sunrise (RSP10989)
- 0.319 Italian Kiss (RSP11034)
- 0.319 RKM-2018-023 (RSP11115)
- 0.321 RKM-2018-027 (RSP11119)
- 0.337 Queen Jesus (RSP10105)
- 0.338 Kimbo Slice (RSP10997)
- 0.340 Skunk 18 (RSP11038)
- 0.348 Gold Cracker (RSP11048)
- 0.350 Cbot-2019-001 (RSP11129)
- 0.350 Tygra (RSP10667)
- 0.352 RKM-2018-031 (RSP11123)
- 0.352 Durban Poison (RSP11014)
- 0.352 Black Beauty (RSP11035)
- 0.356 Blueberry Cheesecake (RSP10684)
- 0.359 Jiangji (RSP10653)
Most genetically distant strains (All Samples)
- 0.525 GMO x Poison Momosa (RSP12626)
- 0.523 Purple Urkle (RSP12890)
- 0.515 JL Cross 14 (RSP11515)
- 0.514 JL Cross 6 (RSP11507)
- 0.499 CS (RSP11208)
- 0.497 BagSeed (RSP12627)
- 0.496 Cherry Blossom (RSP11335)
- 0.495 Cherry Blossom (RSP11308)
- 0.492 CHEM4 (RSP12090)
- 0.492 Deadhead OG (RSP11463)
- 0.491 80E (RSP11213)
- 0.491 Fatso (RSP11741)
- 0.489 Red Eye OG (RSP11190)
- 0.489 Cherry Blossom (RSP11333)
- 0.486 Feral (RSP11205)
- 0.483 Wedding Cake x MAC (RSP11464)
- 0.480 80E (RSP11211)
- 0.479 JL Cross 11 (RSP11512)
- 0.479 Carmaleonte (RSP11207)
- 0.478 Cherry Blossom (RSP11311)
Most genetically distant strains (Base Tree)
- 0.468 Cbot-2019-005 (RSP11133)
- 0.447 RKM-2018-002 (RSP11093)
- 0.437 Cherry (RSP11142)
- 0.429 Skywalker OG (RSP10837)
- 0.429 Kush Hemp E1 (RSP11128)
- 0.429 RKM-2018-026 (RSP11118)
- 0.426 RKM-2018-019 (RSP11111)
- 0.424 Cherry (RSP11143)
- 0.414 RKM-2018-034 (RSP11126)
- 0.406 RKM-2018-032 (RSP11124)
- 0.400 USO 31 (RSP10981)
- 0.398 Carmagnola (RSP11037)
- 0.397 Feral (RSP10890)
- 0.397 Monoica (RSP10241)
- 0.397 RKM-2018-022 (RSP11114)
- 0.392 RKM-2018-028 (RSP11120)
- 0.392 Futura 75 (RSP10664)
- 0.389 Fedora 17 (RSP10661)
- 0.386 Kyrgyz Gold (RSP11054)
- 0.386 Cbot-2019-004 (RSP11132)
Nearest genetic relative in Phylos dataset
- Overlapping SNPs:
- 88
- Concordance:
- 51
Nearest genetic relative in Lynch dataset
- Overlapping SNPs:
- 1
- Concordance:
- 1
Blockchain Registration Information
- Transaction ID
-
37b6fd73d1d293a2
a3ab86a1a62ce0b0 bf8809e202e498fd f015f5a63e2cc639 - Stamping Certificate
- Download PDF (39.4 KB)
- SHASUM Hash
-
a37faaca7ab23fc46193706f2490cab2 7724fc1a4f009420 7286b064ae486a3f