PG
RSP 12928
Grower: Zamir Punja
General Information
- Sample Name
- PG - roots from veg plant
- Accession Date
- September 27, 2023
- Reported Plant Sex
- Female
- Report Type
- Whole-Genome Sequencing
- Microbiome
- Krona Plot
- Fungal Microbiome
- Krona Plot
- Bacterial Microbiome
- Krona Plot
- Viral Microbiome
- Krona Plot
The strain rarity visualization shows how distant the strain is from the other cultivars in the Kannapedia database. The y-axis represents genetic distance, getting farther as you go up. The width of the visualization at any position along the y-axis shows how many strains there are in the database at that genetic distance. So, a common strain will have a more bottom-heavy shape, while uncommon and rare cultivars will have a visualization that is generally shifted towards the top.
Chemical Information
Cannabinoid and terpenoid information provided by the grower.
Cannabinoids
No information provided.
Terpenoids
No information provided.
Genetic Information
- Plant Type
- Type II
File Downloads
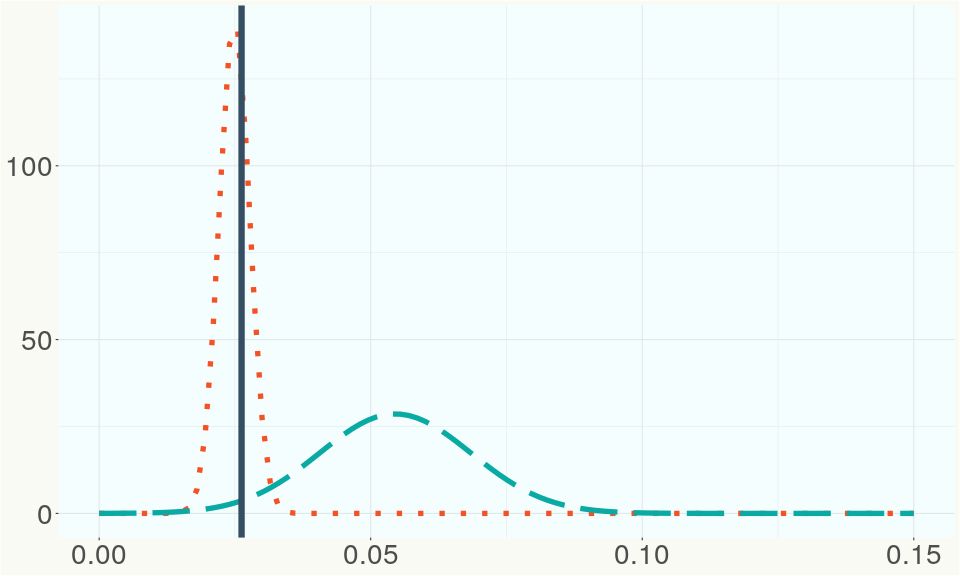
The bell curve in the heterozygosity visualization shows the distribution of heterozygosity levels for cannabis cultivars in the Kannapedia database. The green line shows where this particular strain fits within the distribution. Heterozygosity is associated with heterosis (aka hybrid vigor) but also leads to the production of more variable offspring. When plants have two genetically different parents, heterozygosity levels will be higher than if it has been inbred or backcrossed repeatedly.
The ratio of reads mapped to Y-contigs to reads mapped to the whole Cannabis genome (Y-ratios) has been demonstrated to be strongly correlated with plant sex typing. This plot shows the distribution of Y-ratios for all samples in our database which were sequenced with the same method (panel or WGS) as this sample and where this sample falls in the distribution.
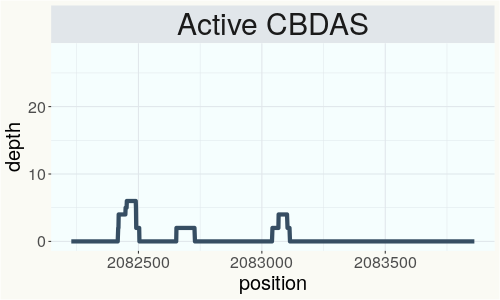
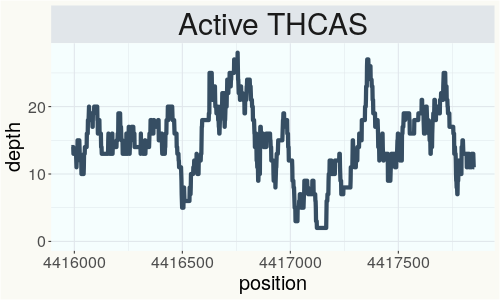
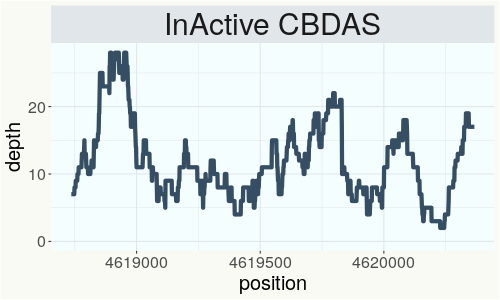
This chart represents the Illumina sequence coverage over the Bt/Bd allele. These are the three regions in the cannabis genome that impact THCA, CBDA, CBGA production. Coverage over the Active CBDAS gene is highly correlated with Type II and Type III plants as described by Etienne de Meijer. Coverage over the THCA gene is highly correlated with Type I and Type II plants but is anti-correlated with Type III plants. Type I plants require coverage over the inactive CBDA loci and no coverage over the Active CBDA gene. Lack of coverage over the Active CBDA and Active THCA allele are presumed to be Type IV plants (CBGA dominant). While deletions of entire THCAS and CBDAS genes are the most common Bt:Bd alleles observed, it is possible to have plants with these genes where functional expression of the enzyme is disrupted by deactivating point mutations (Kojoma et al. 2006).
This chart represents the Illumina sequence coverage over the CBCA synthase gene.
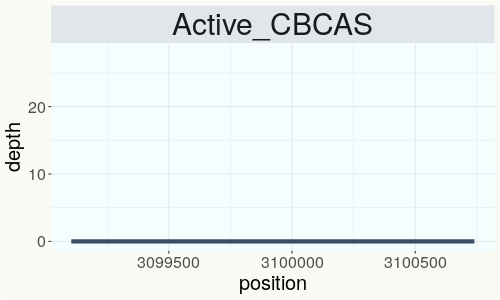
Variants (THCAS, CBDAS, and CBCAS)
Variants (Select Genes of Interest)
| EMF1-2 | c.710A>C | p.His237Pro | missense variant | moderate | contig885 | 810 | A/C | |
| PKSG-2a | c.948T>G | p.Asp316Glu | missense variant | moderate | contig700 | 1944442 | A/C |
|
| PKSG-2a | c.945T>G | p.Ser315Arg | missense variant | moderate | contig700 | 1944445 | A/C |
|
| PKSG-2a | c.944G>A | p.Ser315Asn | missense variant | moderate | contig700 | 1944446 | C/T |
|
| PKSG-2a | c.934C>G | p.His312Asp | missense variant | moderate | contig700 | 1944456 | G/C |
|
| PKSG-2b | c.31A>T | p.Thr11Ser | missense variant | moderate | contig700 | 1951851 | T/A | |
| PKSG-4b |
c.535_545del |
p.Ile179fs | frameshift variant | high | contig700 | 2721127 |
CCCCACTCCAAT |
|
| PKSG-4b | c.496A>G | p.Lys166Glu | missense variant | moderate | contig700 | 2721177 | T/C | |
| PKSG-4b | c.489delT | p.Phe163fs | frameshift variant | high | contig700 | 2721183 | CA/C | |
| PKSG-4b | c.485A>G | p.Lys162Arg | missense variant | moderate | contig700 | 2721188 | T/C | |
| PKSG-4b | c.431T>G | p.Val144Gly | missense variant | moderate | contig700 | 2721242 | A/C | |
| PKSG-4b | c.419A>G | p.Asp140Gly | missense variant | moderate | contig700 | 2721254 | T/C | |
| PKSG-4b | c.316+2T>A | splice donor variant & intron variant | high | contig700 | 2723818 | A/T | ||
| ELF3 | c.358G>A | p.Gly120Arg | missense variant | moderate | contig97 | 242064 | G/A | |
| ELF3 | c.574A>G | p.Asn192Asp | missense variant | moderate | contig97 | 242280 | A/G | |
| ELF3 | c.772A>G | p.Ser258Gly | missense variant | moderate | contig97 | 242478 | A/G |
|
| ELF3 | c.812G>C | p.Gly271Ala | missense variant | moderate | contig97 | 242518 | G/C | |
| ELF3 | c.2140C>T | p.Pro714Ser | missense variant | moderate | contig97 | 244971 | C/T | |
| ELF3 | c.2141C>G | p.Pro714Arg | missense variant | moderate | contig97 | 244972 | C/G | |
| ELF3 | c.2198G>T | p.Arg733Leu | missense variant | moderate | contig97 | 245029 | G/T | |
| aPT4 | c.16G>A | p.Val6Ile | missense variant | moderate | contig121 | 2828672 | G/A |
|
| aPT4 | c.35A>C | p.Gln12Pro | missense variant | moderate | contig121 | 2828691 | A/C |
|
| aPT4 | c.216A>T | p.Lys72Asn | missense variant | moderate | contig121 | 2828872 | A/T |
|
| aPT1 | c.629C>T | p.Thr210Ile | missense variant | moderate | contig121 | 2840237 | C/T | |
| AAE1-2 | c.311A>G | p.Asn104Ser | missense variant | moderate | contig81 | 209273 | A/G |
|
| AAE1-2 | c.331A>G | p.Asn111Asp | missense variant | moderate | contig81 | 209293 | A/G |
|
| AAE1-2 | c.374A>G | p.His125Arg | missense variant | moderate | contig81 | 209336 | A/G |
|
| AAE1-2 |
c.948_949ins |
p.Asp317fs | frameshift variant | high | contig81 | 209910 | C/CA |
|
| AAE1-2 | c.1090A>G | p.Lys364Glu | missense variant | moderate | contig81 | 210052 | A/G |
|
| FAD4 | c.121G>T | p.Val41Phe | missense variant | moderate | contig784 | 1690873 | G/T |
|
| HDS-1 | c.1618A>G | p.Ile540Val | missense variant | moderate | contig1891 | 885936 | T/C | |
| HDS-1 | c.56C>G | p.Ala19Gly | missense variant | moderate | contig1891 | 889336 | G/C | |
| HDS-1 | c.35G>A | p.Cys12Tyr | missense variant | moderate | contig1891 | 889357 | C/T | |
| HDS-1 |
c.-108+1_-10 |
splice donor variant & intron variant | high | contig1891 | 889975 | A/AC |
|
|
| FLD | c.2929T>C | p.Phe977Leu | missense variant | moderate | contig1450 | 2044103 | A/G | |
| GGR | c.362A>T | p.Tyr121Phe | missense variant | moderate | contig2282 | 549354 | A/T |
|
| GGR | c.456T>A | p.His152Gln | missense variant | moderate | contig2282 | 549448 | T/A |
|
| GGR | c.460G>A | p.Asp154Asn | missense variant | moderate | contig2282 | 549452 | G/A |
|
| GGR | c.679G>A | p.Glu227Lys | missense variant | moderate | contig2282 | 549671 | G/A |
|
| GGR | c.704A>T | p.His235Leu | missense variant | moderate | contig2282 | 549696 | A/T |
|
| PKSB-3 | c.1652A>G | p.Glu551Gly | missense variant | moderate | contig93 | 3339759 | A/G |
|
Nearest genetic relatives (All Samples)
- 0.062 PG (RSP12934)
- 0.219 JL Cross 1 (RSP11502)
- 0.222 JFG (RSP12927)
- 0.225 Powdered Donuts (RSP12939)
- 0.230 JFG (RSP12933)
- 0.233 BF (RSP12931)
- 0.242 Old Family Purple (RSP12098)
- 0.247 RKM-2018-010 (RSP11101)
- 0.248 Sunday Driver (RSP11071)
- 0.248 Center Mark (RSP11629)
- 0.250 JL Cross 13 (RSP11514)
- 0.250 Hermaphrodite Research Sample1 (RSP11042)
- 0.251 T S A G E (RSP11351)
- 0.251 Blue Dream (RSP11010)
- 0.252 Blue Dream (RSP11017)
- 0.252 D1 (RSP12875)
- 0.253 BF (RSP12937)
- 0.254 El Gordo (RSP12938)
- 0.255 Blue Dream (RSP11007)
- 0.255 Blue Dream (RSP11004)
Nearest genetic relatives (Base Tree)
- 0.256 Hermaphrodite Research Sample1 (RSP11049)
- 0.272 UP Sunrise (RSP10989)
- 0.277 Blue Dream (RSP11033)
- 0.278 RKM-2018-018 (RSP11110)
- 0.279 RKM-2018-006 (RSP11097)
- 0.281 Blueberry Cheesecake (RSP10684)
- 0.282 Hermaphrodite ResearchSample2 (RSP11050)
- 0.283 Italian Kiss (RSP11034)
- 0.285 RKM-2018-009 (RSP11100)
- 0.287 RKM-2018-028 (RSP11120)
- 0.289 Sour Raspberry (RSP10551)
- 0.297 CST (RSP11002)
- 0.298 Liberty Haze (RSP11000)
- 0.299 RKM-2018-033 (RSP11125)
- 0.306 Gold Cracker (RSP11048)
- 0.310 RKM-2018-003 (RSP11094)
- 0.314 Queen Jesus (RSP10105)
- 0.314 Blueberry Cheesecake (RSP10680)
- 0.315 Pie Hoe (RSP11073)
- 0.315 Cbot-2019-004 (RSP11132)
Most genetically distant strains (All Samples)
- 0.466 Cherry Blossom (RSP11311)
- 0.466 Cherry Blossom (RSP11317)
- 0.457 Cherry Blossom (RSP11328)
- 0.453 Cherry Blossom (RSP11335)
- 0.453 Cherry Blossom (RSP11308)
- 0.451 Cherry Blossom (RSP11334)
- 0.444 Cherry Blossom (RSP11314)
- 0.440 Cherry Blossom (RSP11298)
- 0.431 Feral (RSP11205)
- 0.429 Cherry Blossom (RSP11330)
- 0.429 CS Indica (RSP11658)
- 0.428 CS (RSP11208)
- 0.428 Cherry Blossom (RSP11323)
- 0.428 Cherry Blossom (RSP11312)
- 0.427 Candy Kush (RSP11492)
- 0.426 Unknown- Cherry Wine - 003 (RSP11270)
- 0.425 Cherry Blossom (RSP11333)
- 0.423 HM (RSP12940)
- 0.422 Cherry Blossom (RSP11324)
- 0.420 Cherry Blossom (RSP11309)
Most genetically distant strains (Base Tree)
- 0.405 KYRG-11 (RSP11051)
- 0.403 USO 31 (RSP10981)
- 0.400 Monoica (RSP10241)
- 0.400 Feral (RSP10890)
- 0.395 Futura 75 (RSP10664)
- 0.391 Carmagnola (RSP11037)
- 0.389 Fedora 17 (RSP10661)
- 0.386 Kyrgyz Gold (RSP11054)
- 0.385 Kush Hemp E1 (RSP11128)
- 0.384 Tygra (RSP10667)
- 0.382 Lovrin (RSP10658)
- 0.382 Cbot-2019-005 (RSP11133)
- 0.382 Cherry (RSP11142)
- 0.376 Santhica27 (RSP11047)
- 0.374 Cherry (RSP11143)
- 0.374 Carmagnola (RSP10979)
- 0.373 Tisza (RSP11044)
- 0.367 RKM-2018-026 (RSP11118)
- 0.360 Ivory (RSP10668)
- 0.357 RKM-2018-022 (RSP11114)
Nearest genetic relative in Phylos dataset
- Overlapping SNPs:
- 95
- Concordance:
- 62
Nearest genetic relative in Lynch dataset
- Overlapping SNPs:
- 6
- Concordance:
- 6
Blockchain Registration Information
- Transaction ID
-
1e0458f29d7505e5
70cf4dca42f474b0 a40b5f5b90f56838 a597b0854b99481a - Stamping Certificate
- Download PDF (39.5 KB)
- SHASUM Hash
-
fa7ec322c9b1f89b676d0bc31a9b64cd 922e62ddcfa13011 2800b4f8c58b63be