UC_18
SRR 14419894
General Information
- Accession Date
- April 18, 2021
- Reported Plant Sex
- not reported
- Report Type
- Whole-Genome Sequencing
The strain rarity visualization shows how distant the strain is from the other cultivars in the Kannapedia database. The y-axis represents genetic distance, getting farther as you go up. The width of the visualization at any position along the y-axis shows how many strains there are in the database at that genetic distance. So, a common strain will have a more bottom-heavy shape, while uncommon and rare cultivars will have a visualization that is generally shifted towards the top.
Chemical Information
Cannabinoid and terpenoid information provided by the grower.
Cannabinoids
- THC + THCA
- 0.0011%
- CBD + CBDA
- 0.0378%
- THCV + THCVA
- n/a
- CBC + CBCA
- 0.%
- CBG + CBGA
- 0.0009%
- CBN + CBNA
- n/a
Terpenoids
- α-Bisabolol
- n/a
- Borneol
- n/a
- Camphene
- 0.0001%
- Carene
- 0.%
- Caryophyllene oxide
- 0.0011%
- β-Caryophyllene
- 0.0014%
- Fenchol
- n/a
- Geraniol
- 0.%
- α-Humulene
- n/a
- Limonene
- n/a
- Linalool
- 0.%
- Myrcene
- n/a
- α-Phellandrene
- n/a
- Terpinolene
- n/a
- α-Terpineol
- n/a
- α-Terpinene
- 0.0002%
- γ-Terpinene
- 0.0001%
- Total Nerolidol
- n/a
- Total Ocimene
- 0.%
- α-Pinene
- 0.0043%
- β-Pinene
- n/a
Genetic Information
- Plant Type
- Type III
File Downloads
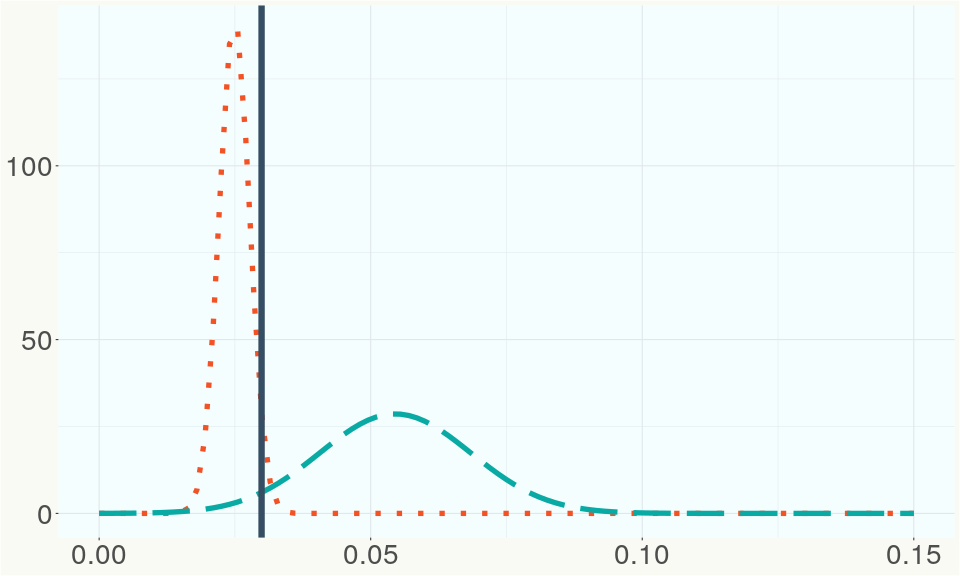
The bell curve in the heterozygosity visualization shows the distribution of heterozygosity levels for cannabis cultivars in the Kannapedia database. The green line shows where this particular strain fits within the distribution. Heterozygosity is associated with heterosis (aka hybrid vigor) but also leads to the production of more variable offspring. When plants have two genetically different parents, heterozygosity levels will be higher than if it has been inbred or backcrossed repeatedly.
The ratio of reads mapped to Y-contigs to reads mapped to the whole Cannabis genome (Y-ratios) has been demonstrated to be strongly correlated with plant sex typing. This plot shows the distribution of Y-ratios for all samples in our database which were sequenced with the same method (panel or WGS) as this sample and where this sample falls in the distribution.
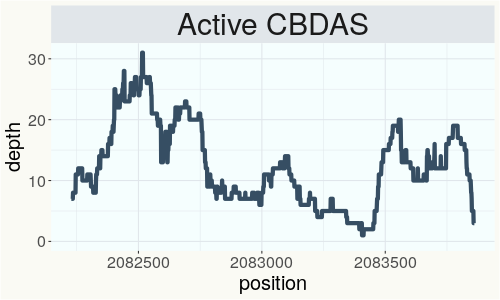
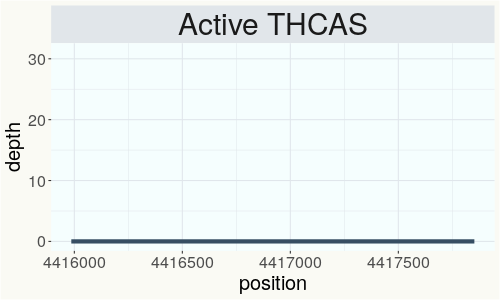
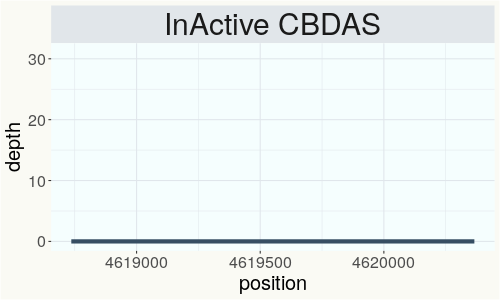
This chart represents the Illumina sequence coverage over the Bt/Bd allele. These are the three regions in the cannabis genome that impact THCA, CBDA, CBGA production. Coverage over the Active CBDAS gene is highly correlated with Type II and Type III plants as described by Etienne de Meijer. Coverage over the THCA gene is highly correlated with Type I and Type II plants but is anti-correlated with Type III plants. Type I plants require coverage over the inactive CBDA loci and no coverage over the Active CBDA gene. Lack of coverage over the Active CBDA and Active THCA allele are presumed to be Type IV plants (CBGA dominant). While deletions of entire THCAS and CBDAS genes are the most common Bt:Bd alleles observed, it is possible to have plants with these genes where functional expression of the enzyme is disrupted by deactivating point mutations (Kojoma et al. 2006).
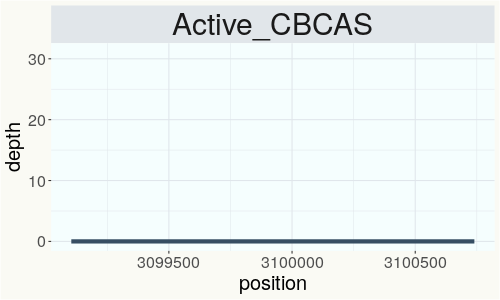
This chart represents the Illumina sequence coverage over the CBCA synthase gene.
Variants (THCAS, CBDAS, and CBCAS)
No variants to report
Variants (Select Genes of Interest)
| EMF1-2 | c.1187T>C | p.Leu396Ser | missense variant | moderate | contig885 | 2073 | T/C |
|
| PHL-2 | c.932T>C | p.Leu311Pro | missense variant | moderate | contig2621 | 340210 | T/C | |
| PHL-2 | c.1057A>G | p.Arg353Gly | missense variant | moderate | contig2621 | 340335 | A/G | |
| PHL-2 | c.2564T>A | p.Phe855Tyr | missense variant | moderate | contig2621 | 342607 | T/A | |
| PHL-2 | c.2578T>A | p.Leu860Ile | missense variant | moderate | contig2621 | 342621 | T/A | |
| PHL-2 | c.2624C>T | p.Ser875Phe | missense variant | moderate | contig2621 | 342667 | C/T | |
| PHL-2 | c.2783G>A | p.Ser928Asn | missense variant | moderate | contig2621 | 342826 | G/A | |
| PHL-2 | c.2830A>C | p.Asn944His | missense variant | moderate | contig2621 | 342873 | A/C |
|
| PHL-2 | c.2933G>T | p.Arg978Leu | missense variant | moderate | contig2621 | 342976 | G/T | |
| PHL-2 | c.2936T>G | p.Val979Gly | missense variant | moderate | contig2621 | 342979 | T/G |
|
| PHL-2 | c.3209A>G | p.Gln1070Arg | missense variant | moderate | contig2621 | 343252 | A/G | |
| PHL-2 | c.3467A>G | p.Gln1156Arg | missense variant | moderate | contig2621 | 343510 | A/G | |
| PKSG-4b |
c.535_545del |
p.Ile179fs | frameshift variant | high | contig700 | 2721127 |
CCCCACTCCAAT |
|
| PKSG-4b | c.523C>T | p.His175Tyr | missense variant | moderate | contig700 | 2721150 | G/A | |
| PKSG-4b | c.496A>G | p.Lys166Glu | missense variant | moderate | contig700 | 2721177 | T/C | |
| PKSG-4b | c.489delT | p.Phe163fs | frameshift variant | high | contig700 | 2721183 | CA/C | |
| PKSG-4b | c.485A>G | p.Lys162Arg | missense variant | moderate | contig700 | 2721188 | T/C | |
| PKSG-4b | c.431T>G | p.Val144Gly | missense variant | moderate | contig700 | 2721242 | A/C | |
| PKSG-4b |
c.352_355del |
p.Thr118fs | frameshift variant | high | contig700 | 2721317 | CCTGT/C |
|
| PKSG-4b |
c.353_354ins |
p.Gly119fs | frameshift variant | high | contig700 | 2721319 | T/TGG |
|
| PKSG-4b | c.323A>G | p.Glu108Gly | missense variant | moderate | contig700 | 2721350 | T/C |
|
| PKSG-4b | c.316+2T>A | splice donor variant & intron variant | high | contig700 | 2723818 | A/T | ||
| PKSG-4b | c.238T>C | p.Phe80Leu | missense variant | moderate | contig700 | 2724197 | A/G |
|
| PKSG-4b | c.229G>A | p.Gly77Ser | missense variant | moderate | contig700 | 2724206 | C/T |
|
| PKSG-4b | c.216G>C | p.Leu72Phe | missense variant | moderate | contig700 | 2724219 | C/G |
|
| PKSG-4b | c.206T>C | p.Leu69Ser | missense variant | moderate | contig700 | 2724229 | A/G |
|
| OAC-1 | c.260C>G | p.Ser87Cys | missense variant | moderate | contig931 | 118104 | G/C |
|
| OAC-1 | c.185C>T | p.Thr62Ile | missense variant | moderate | contig931 | 118179 | G/A |
|
| OAC-1 | c.175G>A | p.Val59Ile | missense variant | moderate | contig931 | 118189 | C/T |
|
| FAD2-2 | c.161T>A | p.Leu54His | missense variant | moderate | contig83 | 1803208 | A/T |
|
| FAD2-2 | c.127C>T | p.Arg43* | stop gained | high | contig83 | 1803242 | G/A |
|
| FAD2-2 | c.75C>A | p.His25Gln | missense variant | moderate | contig83 | 1803294 | G/T |
|
| ELF3 |
c.364_366del |
p.Lys122del | conservative inframe deletion | moderate | contig97 | 242066 | AAAG/A |
|
| ELF3 | c.772A>G | p.Ser258Gly | missense variant | moderate | contig97 | 242478 | A/G |
|
| ELF3 | c.812G>C | p.Gly271Ala | missense variant | moderate | contig97 | 242518 | G/C | |
| ELF3 | c.1366T>G | p.Leu456Val | missense variant | moderate | contig97 | 244197 | T/G |
|
| ELF3 | c.1435G>C | p.Ala479Pro | missense variant | moderate | contig97 | 244266 | G/C |
|
| ELF3 | c.1966C>G | p.Pro656Ala | missense variant | moderate | contig97 | 244797 | C/G | |
| ELF3 | c.2198G>T | p.Arg733Leu | missense variant | moderate | contig97 | 245029 | G/T | |
| HDS-2 | c.679G>C | p.Gly227Arg | missense variant | moderate | contig95 | 1990632 | G/C |
|
| AAE1-2 | c.331A>G | p.Asn111Asp | missense variant | moderate | contig81 | 209293 | A/G |
|
| AAE1-2 | c.409C>T | p.Arg137Cys | missense variant | moderate | contig81 | 209371 | C/T |
|
| AAE1-2 | c.1006A>G | p.Lys336Glu | missense variant | moderate | contig81 | 209968 | A/G |
|
| AAE1-2 | c.1102C>A | p.His368Asn | missense variant | moderate | contig81 | 210064 | C/A |
|
| AAE1-2 | c.1541T>C | p.Val514Ala | missense variant | moderate | contig81 | 210503 | T/C |
|
| PHL-1 | c.2623A>G | p.Thr875Ala | missense variant | moderate | contig1439 | 1487174 | T/C |
|
| PHL-1 | c.2551A>G | p.Thr851Ala | missense variant | moderate | contig1439 | 1487246 | T/C | |
| PHL-1 | c.1387A>G | p.Thr463Ala | missense variant | moderate | contig1439 | 1489811 | T/C | |
| Edestin | c.40G>A | p.Ala14Thr | missense variant | moderate | contig850 | 3065250 | C/T |
|
| Edestin | c.17C>T | p.Ser6Leu | missense variant | moderate | contig850 | 3065273 | G/A |
|
| Edestin | c.16T>C | p.Ser6Pro | missense variant | moderate | contig850 | 3065274 | A/G |
|
| HDS-1 | c.1378G>A | p.Val460Ile | missense variant | moderate | contig1891 | 886370 | C/T |
|
| PIE1-2 | c.6653A>G | p.Asn2218Ser | missense variant | moderate | contig1460 | 1184434 | T/C | |
| PIE1-2 | c.2188G>A | p.Ala730Thr | missense variant | moderate | contig1460 | 1189852 | C/T | |
| PIE1-2 | c.1872T>A | p.Asp624Glu | missense variant | moderate | contig1460 | 1190252 | A/T | |
| EMF2 | c.1772A>G | p.Gln591Arg | missense variant | moderate | contig954 | 3059929 | A/G | |
| EMF1-1 | c.242A>G | p.Lys81Arg | missense variant | moderate | contig883 | 269731 | A/G | |
| FT |
c.413_413+1i |
splice donor variant & intron variant | high | contig1561 | 3126450 | T/TGTATGCA |
|
|
| AAE1-3 | c.667G>A | p.Val223Ile | missense variant | moderate | contig976 | 1083187 | C/T |
|
| AAE1-3 | c.655C>T | p.Pro219Ser | missense variant | moderate | contig976 | 1083199 | G/A |
|
| AAE1-3 | c.635G>A | p.Gly212Asp | missense variant | moderate | contig976 | 1083219 | C/T |
|
| AAE1-3 | c.634G>C | p.Gly212Arg | missense variant | moderate | contig976 | 1083220 | C/G |
|
| AAE1-3 | c.403G>T | p.Asp135Tyr | missense variant | moderate | contig976 | 1083622 | C/A |
|
| AAE1-3 | c.382T>C | p.Tyr128His | missense variant | moderate | contig976 | 1083643 | A/G |
|
| AAE1-3 | c.367C>T | p.Arg123Trp | missense variant | moderate | contig976 | 1083658 | G/A |
|
| AAE1-3 | c.296C>T | p.Pro99Leu | missense variant | moderate | contig976 | 1083729 | G/A |
|
| AAE1-3 | c.293A>G | p.Asp98Gly | missense variant | moderate | contig976 | 1083732 | T/C |
|
| AAE1-3 | c.284A>T | p.Glu95Val | missense variant | moderate | contig976 | 1083741 | T/A |
|
| AAE1-3 | c.267T>G | p.His89Gln | missense variant | moderate | contig976 | 1083758 | A/C |
|
| AAE1-3 | c.224C>T | p.Pro75Leu | missense variant | moderate | contig976 | 1083851 | G/A |
|
| AAE1-3 | c.208G>C | p.Ala70Pro | missense variant | moderate | contig976 | 1083867 | C/G |
|
| AAE1-3 | c.181G>A | p.Val61Ile | missense variant | moderate | contig976 | 1083894 | C/T |
|
| AAE1-3 | c.167A>G | p.Glu56Gly | missense variant | moderate | contig976 | 1083908 | T/C |
|
| AAE1-3 | c.154G>A | p.Val52Ile | missense variant | moderate | contig976 | 1083921 | C/T |
|
| AAE1-3 | c.125A>G | p.Glu42Gly | missense variant | moderate | contig976 | 1083950 | T/C |
|
| AAE1-3 | c.101A>G | p.Tyr34Cys | missense variant | moderate | contig976 | 1083974 | T/C |
|
| AAE1-3 | c.79A>G | p.Thr27Ala | missense variant | moderate | contig976 | 1083996 | T/C |
|
| AAE1-3 | c.52G>A | p.Gly18Ser | missense variant | moderate | contig976 | 1084023 | C/T |
|
| AAE1-3 | c.12G>A | p.Met4Ile | missense variant | moderate | contig976 | 1084063 | C/T |
|
| AAE1-3 | c.11T>C | p.Met4Thr | missense variant | moderate | contig976 | 1084064 | A/G |
|
| AAE1-3 | c.3G>A | p.Met1? | start lost | high | contig976 | 1084072 | C/T | |
| PIE1-1 | c.1222C>G | p.Gln408Glu | missense variant | moderate | contig1225 | 2281482 | C/G | |
| GGR | c.376G>C | p.Glu126Gln | missense variant | moderate | contig2282 | 549368 | G/C | |
| GGR | c.382C>T | p.Leu128Phe | missense variant | moderate | contig2282 | 549374 | C/T |
|
| GGR | c.456T>A | p.His152Gln | missense variant | moderate | contig2282 | 549448 | T/A |
|
| GGR | c.460G>A | p.Asp154Asn | missense variant | moderate | contig2282 | 549452 | G/A |
|
| GGR | c.581C>T | p.Ala194Val | missense variant | moderate | contig2282 | 549573 | C/T | |
| GGR | c.704A>T | p.His235Leu | missense variant | moderate | contig2282 | 549696 | A/T |
|
| PKSB-3 | c.1490C>T | p.Ala497Val | missense variant | moderate | contig93 | 3339597 | C/T |
|
| PKSB-3 | c.1652A>G | p.Glu551Gly | missense variant | moderate | contig93 | 3339759 | A/G |
|
| PKSB-3 | c.1848G>A | p.Met616Ile | missense variant | moderate | contig93 | 3339955 | G/A |
|
Nearest genetic relatives (All Samples)
- 0.052 PCL1 (SRR14708246)
- 0.088 VIR 37 - Novgorod-Seversky - cv (SRR14708234)
- 0.089 Tiborszallasi (SRR14708273)
- 0.094 VIR 223 - Bernburgskaya Odnodomnaya - bm (SRR14708217)
- 0.100 VIR 483 (SRR14708238)
- 0.100 PCL2 (SRR14708245)
- 0.107 Triangle Kush x Square Wave BX (RSP12100)
- 0.107 Carmagnola (SRR14708200)
- 0.111 Mexican Sativa (SRR14708263)
- 0.113 R2in135 (SRR14708236)
- 0.116 UC 32 (SRR14419761)
- 0.118 VIR 483 (SRR14708240)
- 0.121 UC 167 (SRR14419585)
- 0.121 Electra (RSP11366)
- 0.121 Kompolti (SRR14708277)
- 0.122 XUM2 (SRR14708204)
- 0.122 Wild Thailand (SRR14708216)
- 0.125 VIR 369 (SRR14708231)
- 0.125 Fibranova (SRR14708276)
- 0.126 UC 46 (SRR14419703)
Most genetically distant strains (All Samples)
- 0.412 80E (RSP11213)
- 0.387 JL yellow (RSP11075)
- 0.378 80E (RSP11211)
- 0.374 JL 3rd Gen Mother (RSP11214)
- 0.374 Skunk 18 (RSP11038)
- 0.373 JL 4th Gen 5 (RSP11199)
- 0.366 Tanao Sri 46 (RSP11486)
- 0.360 Cherry Blossom (RSP11318)
- 0.359 80E (RSP11212)
- 0.359 Chem 91 (RSP11185)
- 0.357 JL 4th Gen 2 (RSP11194)
- 0.353 JL 3rd Gen Father (RSP11196)
- 0.350 Northern Lights (RSP11501)
- 0.350 Tanao Sri-white 80 (RSP11621)
- 0.348 Tiger Tail 30 (RSP11484)
- 0.347 Cbot-2019-005 (RSP11133)
- 0.341 White Label 1 (RSP11336)
- 0.340 Kush Hemp E1 (RSP11128)
- 0.340 Cherry Blossom (RSP11323)
- 0.339 Chematonic Cannatonic x Chemdawg (RSP11394)
Nearest genetic relative in Phylos dataset
- Overlapping SNPs:
- 17
- Concordance:
- 15
Blockchain Registration Information
- SHASUM Hash
-
aa5594fb30fc8ff9eeb15ab2487813e5 58b672abe27a9530 8e8b211a2f4db110